Estimating the pi by throwing pebbles on the sand¶
The key idea of this technique is that the ratio of the area of the circle to the square area that inscribes it is , so by counting the fraction of the random points in the square that are inside the circle, we get increasingly good estimates to .
Source
import numpy as np
import matplotlib.pyplot as plt
def circle_pi_estimate(N=10000, r0=1):
"""
Estimate the value of pi using the Monte Carlo method.
Generate N random points in a square with sides ranging from -r0 to r0.
Count the fraction of points that fall inside the inscribed circle to estimate pi.
Parameters:
N (int): Number of points to generate (default: 10000)
r0 (int): Radius of the circle (default: 1)
Returns:
float: Estimated value of pi
"""
# Generate random points
xs = np.random.uniform(-r0, r0, size=N)
ys = np.random.uniform(-r0, r0, size=N)
# Calculate distances from the origin and determine points inside the circle
inside = np.sqrt(xs**2 + ys**2) < r0
# Compute volume ratio as the ratio of points
v_ratio = inside.sum() / N
pi_estimate = 4 * v_ratio
# Plotting
fig, ax = plt.subplots(figsize=(6, 6))
ax.plot(xs[inside], ys[inside], 'b.', label='Inside')
ax.plot(xs[~inside], ys[~inside], 'r.', label='Outside')
ax.set_title(f"Estimation of $\\pi$ = {pi_estimate}", fontsize=20)
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend()
return pi_estimatecircle_pi_estimate(N=100000, r0=100)3.14432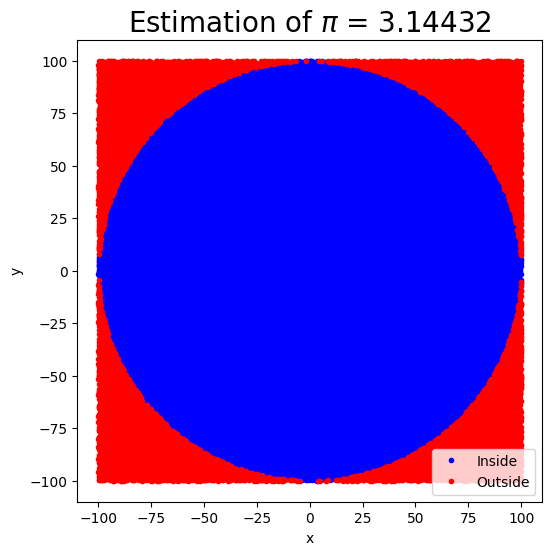
We will now use the same technique but compute a 1D definite integral from to by drawing a rectangle to cover the curve with dimensions and .
The area of the rectangle is . The area under the curve is .
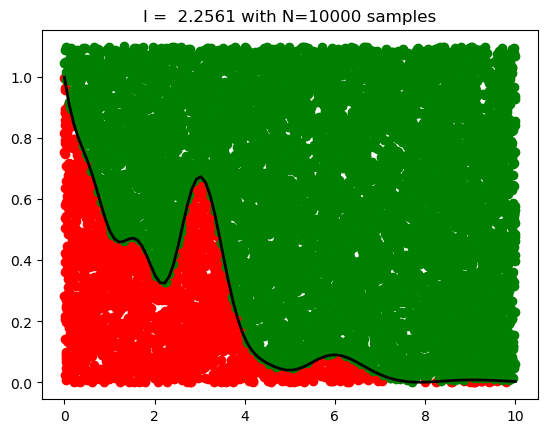
If we choose a point uniformly at random in the rectangle, what is the probability that the point falls into the region under the curve? It is simply
Thus we can estimate the definite integral by drawing uniform points covering the rectangle and computing the integral as
Source
def myfunc1(x):
return np.exp(-x)+ np.exp(-x)* x**2 * np.cos(x)**2 + np.exp(-2*x)*x**4* np.cos(2*x)**2
x= np.linspace(0, 10, 1000)
y = myfunc1(x)
plt.plot(x,y, c='k')
plt.fill_between(x, y,color='gold',alpha=0.3)
plt.xlabel('x')
plt.ylabel('f(x)')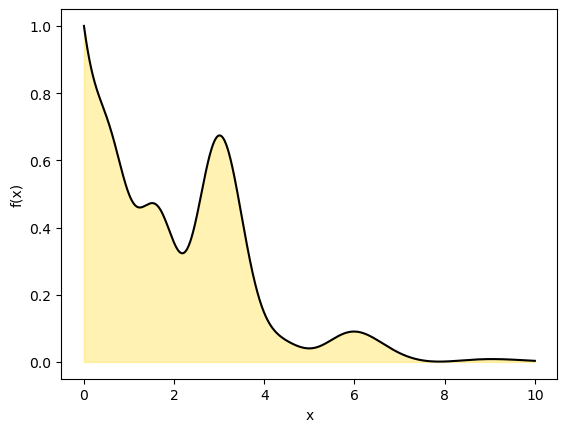
Source
def mc_integral(func,
N=10000,
Lx=2, Ly=1,
plot=True):
'''Generate random points in the square with [0, Lx] and [0, Ly]
Count the fraction of points falling inside the curve
'''
# Generate uniform random numbers
ux = Lx*np.random.rand(N)
uy = Ly*np.random.rand(N)
#Count accepted point.
pinside = uy<func(ux)
# Total area times fraction of sucessful points
I = Lx*Ly*pinside.sum()/N
if plot==True:
plt.plot(ux[pinside], uy[pinside],'o', color='red')
plt.plot(ux[~pinside], uy[~pinside],'o', color='green')
x = np.linspace(0.001, Lx,100)
plt.plot(x, func(x), color='black', lw=2)
plt.title(f'I = {I:.4f} with N={N} samples',fontsize=12)
return I
### Calculate integral numerically first
from scipy import integrate
#adjust limits of x and y
Lx = 10 # x range from 0 to Lx
Ly = 1.1 # y range from 0 to Ly
y, err = integrate.quad(myfunc1, 0, Lx)
print("Exact result:", y)
I = mc_integral(myfunc1, N=10000, Lx=Lx, Ly=Ly)
print("MC result:", I)Exact result: 2.289834301866351
MC result: 2.2561
The Essence of Monte Carlo Simulations¶
Suppose we want to evaluate an integral .
A powerful perspective is to reinterpret this integral as the expectation of a function under some probability distribution :
In this form, the integral becomes the expected value of with respect to the distribution .
To estimate , we draw samples and apply the law of large numbers, which guarantees that the sample average converges to the expected value as :
Simple 1D applications of MC¶
Ordinary Monte Carlo and Uniform Sampling¶
A common and intuitive case is when we draw samples uniformly from the interval . In this setting, the sampling distribution is constant: , and the integral simplifies as follows:
This gives a clear interpretation of Monte Carlo integration: we approximate the average height of the function over the interval by randomly sampling points, much like tossing pebbles onto a plot and estimating the shaded area.
Source
x0, x1 = 0, 10
N = 100000
x = np.random.uniform(x0, x1, N)
integral = (x1 - x0) * np.mean(myfunc1(x))
print('MC result', integral)
y, err = integrate.quad(myfunc1, x0, x1)
print("Exact result:", y)MC result 2.312692930908909
Exact result: 2.289834301866351
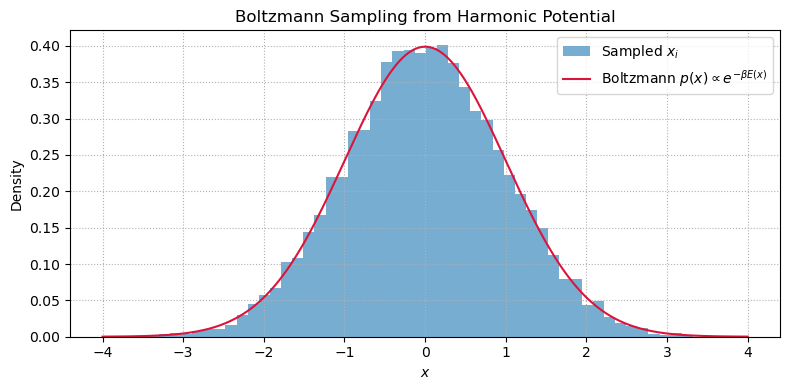
Sampling from the Boltzmann Distribution¶
Quantities like average energy, heat capacity, or pressure are computed as ensemble averages under the Boltzmann distribution.
The Boltzmann distribution for a system with energy at inverse temperature is:
Suppose we are interested in the average energy:
This is an expectation value under the Boltzmann distribution .
If we can draw samples , we can estimate by:
Source
import numpy as np
import matplotlib.pyplot as plt
# Parameters
beta = 1.0 # inverse temperature
n_samples = 10000
# Boltzmann sampling for harmonic oscillator: Gaussian with variance 1/beta
x_samples = np.random.normal(loc=0.0, scale=np.sqrt(1 / beta), size=n_samples)
# Energy function E(x) = (1/2) * x^2
E = 0.5 * x_samples**2
# Estimate average energy ⟨E⟩
E_mean = np.mean(E)
E_exact = 0.5 / beta
print(f"Estimated ⟨E⟩: {E_mean:.4f}")
print(f"Exact ⟨E⟩: {E_exact:.4f}")
# Plot histogram and overlay Boltzmann density
x_vals = np.linspace(-4, 4, 500)
p_vals = np.exp(-0.5 * beta * x_vals**2)
p_vals /= np.trapezoid(p_vals, x_vals) # normalize for display
plt.figure(figsize=(8, 4))
plt.hist(x_samples, bins=50, density=True, alpha=0.6, label='Sampled $x_i$')
plt.plot(x_vals, p_vals, color='crimson', label='Boltzmann $p(x) \\propto e^{-\\beta E(x)}$')
plt.xlabel('$x$')
plt.ylabel('Density')
plt.title('Boltzmann Sampling from Harmonic Potential')
plt.legend()
plt.grid(True, linestyle=':')
plt.tight_layout()
plt.show()Estimated ⟨E⟩: 0.4959
Exact ⟨E⟩: 0.5000
MC Integration in Higher Dimensions¶
Monte Carlo really shines in higher dimensions. Let’s see it in action on a 2D function with sharp features — a mixture of Gaussians:
This is a good proxy for physical problems where the integrand has localized peaks (e.g., partition functions, reaction coordinates).
Notice how most uniform samples miss the peaks entirely — foreshadowing the need for smarter sampling.
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# ── Define a 2D Gaussian mixture ────────────────────────────────
centers = [(-1.5, -1), (1.5, 1.5), (0, -1.5), (-1, 2)]
sigmas = [0.4, 0.5, 0.35, 0.45]
weights = [1.0, 0.7, 0.9, 0.6]
def gmm_2d(x, y):
"""2D Gaussian mixture — a peaked, multi-modal function."""
f = np.zeros_like(x, dtype=float)
for (cx, cy), s, w in zip(centers, sigmas, weights):
f += w * np.exp(-((x - cx)**2 + (y - cy)**2) / (2 * s**2))
return f
# ── MC samples (uniform) ──────────────────────────────────────
N = 5000
xs = np.random.uniform(-4, 4, N)
ys = np.random.uniform(-4, 4, N)
fs = gmm_2d(xs, ys)
# ── Surface grid ───────────────────────────────────────────────
xg = np.linspace(-4, 4, 200)
yg = np.linspace(-4, 4, 200)
X, Y = np.meshgrid(xg, yg)
Z = gmm_2d(X, Y)
# ── Figure ─────────────────────────────────────────────────────
fig = plt.figure(figsize=(16, 6))
ax1 = fig.add_subplot(121, projection='3d')
ax1.plot_surface(X, Y, Z, cmap='viridis', alpha=0.7, edgecolor='none')
ax1.scatter(xs[::3], ys[::3], fs[::3], c=fs[::3], cmap='hot', s=4, alpha=0.6)
ax1.set(xlabel='$x$', ylabel='$y$', zlabel='$f(x,y)$')
ax1.set_title('MC samples on 2D Gaussian mixture', fontsize=13)
ax1.view_init(elev=30, azim=-45)
ax2 = fig.add_subplot(122)
cont = ax2.contourf(X, Y, Z, levels=30, cmap='viridis', alpha=0.5)
ax2.scatter(xs, ys, c=fs, cmap='hot', s=6, alpha=0.7, edgecolors='none')
plt.colorbar(cont, ax=ax2, label='$f(x,y)$')
for (cx, cy) in centers:
ax2.plot(cx, cy, 'w*', markersize=12, markeredgecolor='k', markeredgewidth=0.6)
ax2.set(xlabel='$x$', ylabel='$y$', aspect='equal')
ax2.set_title('Top view: most uniform samples miss the peaks', fontsize=13)
plt.tight_layout()
plt.show()
area = 8 * 8
I_mc = area * np.mean(fs)
print(f"MC integral estimate (N={N}): {I_mc:.4f}")
print(f"Fraction of samples where f > 0.1: {np.mean(fs > 0.1):.2%}")MC vs Grid: the Curse of Dimensionality¶
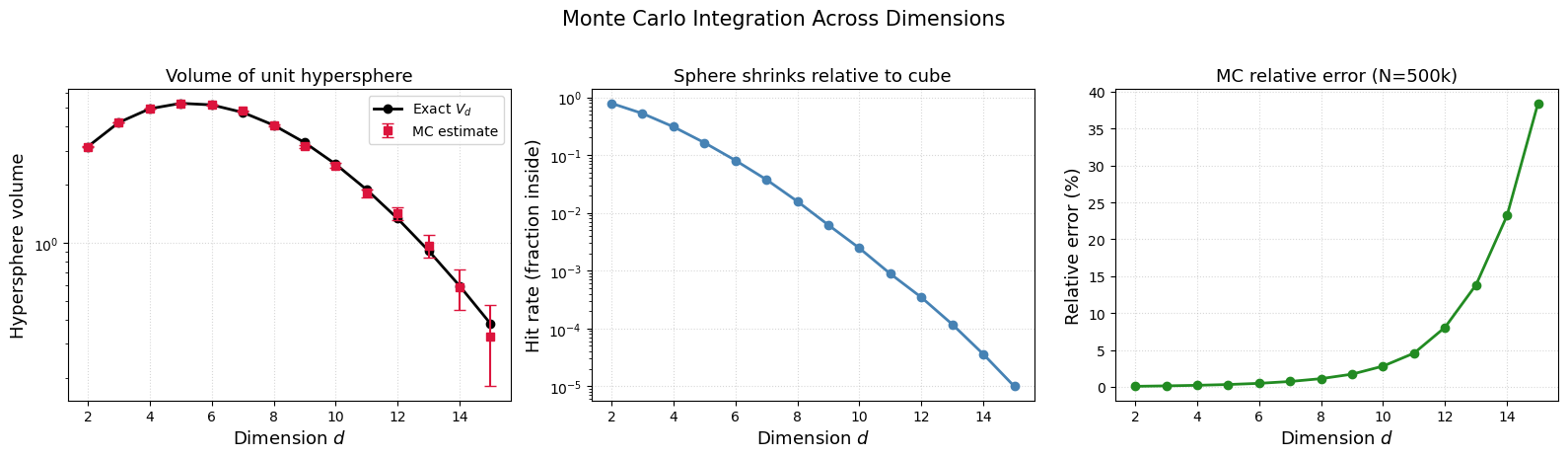
The volume of a unit hypersphere in dimensions is .
We can estimate it by MC: throw uniform points in the hypercube, count the fraction inside , and multiply by the cube volume .
As grows, the sphere shrinks relative to the cube — the “hit rate” drops exponentially.
Two ways to do this MC integral — and they behave differently in high :
| Method | Formula | What happens as |
|---|---|---|
| Hit-or-miss | Hit rate , so grows fails | |
| Mean-value | Same formula, convergence rate is still |
The convergence rate is always dimension-independent — that is the superpower of MC over grid methods (which need points).
But itself can grow with when you sample uniformly in a region where the integrand is mostly zero. This does not mean MC is broken — it means uniform sampling is a bad choice of in high dimensions.
This is exactly what motivates importance sampling (choose a smarter ) and eventually MCMC (let the chain find the important regions automatically).
Source
import numpy as np
import matplotlib.pyplot as plt
from scipy.special import gamma
def hypersphere_volume_exact(d):
"""Exact volume of a unit hypersphere in d dimensions."""
return np.pi**(d / 2) / gamma(d / 2 + 1)
def mc_hypersphere(d, N=500000):
"""MC estimate of the unit hypersphere volume in d dimensions."""
points = np.random.uniform(-1, 1, size=(N, d))
inside = np.sum(points**2, axis=1) < 1.0
hit_rate = np.mean(inside)
cube_vol = 2**d
V_mc = cube_vol * hit_rate
V_err = cube_vol * np.std(inside) / np.sqrt(N)
return V_mc, V_err, hit_rate
dims = np.arange(2, 16)
V_exact = [hypersphere_volume_exact(d) for d in dims]
V_mc_vals, V_err_vals, hit_rates = zip(*[mc_hypersphere(d) for d in dims])
fig, axes = plt.subplots(1, 3, figsize=(16, 4.5))
# Left: exact vs MC volume
axes[0].semilogy(dims, V_exact, 'ko-', label='Exact $V_d$', lw=2)
axes[0].errorbar(dims, V_mc_vals, yerr=V_err_vals, fmt='s', color='crimson',
capsize=4, label='MC estimate')
axes[0].set_xlabel('Dimension $d$', fontsize=13)
axes[0].set_ylabel('Hypersphere volume', fontsize=13)
axes[0].set_title('Volume of unit hypersphere', fontsize=13)
axes[0].legend()
axes[0].grid(True, linestyle=':', alpha=0.5)
# Middle: hit rate (fraction inside sphere)
axes[1].semilogy(dims, hit_rates, 'o-', color='steelblue', lw=2)
axes[1].set_xlabel('Dimension $d$', fontsize=13)
axes[1].set_ylabel('Hit rate (fraction inside)', fontsize=13)
axes[1].set_title('Sphere shrinks relative to cube', fontsize=13)
axes[1].grid(True, linestyle=':', alpha=0.5)
# Right: relative error GROWS with d
rel_err = np.array(V_err_vals) / np.array(V_exact)
axes[2].plot(dims, rel_err * 100, 'o-', color='forestgreen', lw=2)
axes[2].set_xlabel('Dimension $d$', fontsize=13)
axes[2].set_ylabel('Relative error (%)', fontsize=13)
axes[2].set_title(r'Relative error grows: $\sigma$ grows with $d$', fontsize=13)
axes[2].grid(True, linestyle=':', alpha=0.5)
plt.suptitle('Uniform MC integration across dimensions (N=500k)', fontsize=15, y=1.02)
plt.tight_layout()
plt.show()
print(f"\nAt d=2: hit rate = {hit_rates[0]:.3f}, relative error = {rel_err[0]*100:.1f}%")
print(f"At d=15: hit rate = {hit_rates[-1]:.5f}, relative error = {rel_err[-1]*100:.1f}%")
print(f"\nThe rate is always O(1/sqrt(N)), but sigma grows => need smarter sampling!")
Why Does Monte Carlo Outperform Grids? And How to Reduce MC Error¶
By the Central Limit Theorem, the MC estimator has standard error , independent of dimension.
Deterministic grid methods converge as — they slow down exponentially as dimensionality grows (the curse of dimensionality).
The MC error has two independent knobs:
| Knob | What it does | How |
|---|---|---|
| Increase | Shrink | Throw more samples (brute force) |
| Decrease | Shrink the prefactor | Sample where the integrand matters (importance sampling) |
When uniform sampling fails: If is sharply peaked (e.g., a Boltzmann factor at low ), most uniform samples land where . The variance becomes enormous.
Importance sampling attacks directly: by sampling from a distribution that concentrates points where is large, the reweighted values become nearly constant, dramatically reducing variance.
We will vary the sample size from 1 to 100 and calculate the value of for 1000 replicates. We then plot the 2.5th and 97.5th percentile of the 1000 values of to see how the variation in changes with sample size.
def f(x):
return x * np.cos(71*x) + np.sin(13*x)
x = np.linspace(0, 1, 100)
plt.plot(x, f(x), linewidth=2.0)
plt.xlabel(r'$x$', fontsize=16)
plt.ylabel(r'$f(x)$', fontsize=16);Source
n = 100
reps = 1000
# Generating random numbers and applying the function
fx = f(np.random.random((n, reps)))
# Calculating cumulative mean for each simulation
y = np.cumsum(fx, axis=0) / np.arange(1, n+1)[:, None]
# Calculating the upper and lower percentiles
upper, lower = np.percentile(y, [97.5, 2.5], axis=1)
# Plotting the results
plt.figure(figsize=(10, 6))
for i in range(reps):
plt.plot(np.arange(1, n+1), y[:, i], c='grey', alpha=0.02)
plt.plot(np.arange(1, n+1), y[:, 0], c='red', linewidth=1)
plt.plot(np.arange(1, n+1), upper, 'b', label='97.5th percentile')
plt.plot(np.arange(1, n+1), lower, 'b', label='2.5th percentile')
plt.xlabel('Number of independent MC runs')
plt.ylabel('Cumulative mean of f(x)')
plt.title('Monte Carlo Simulations')
plt.legend()
plt.show()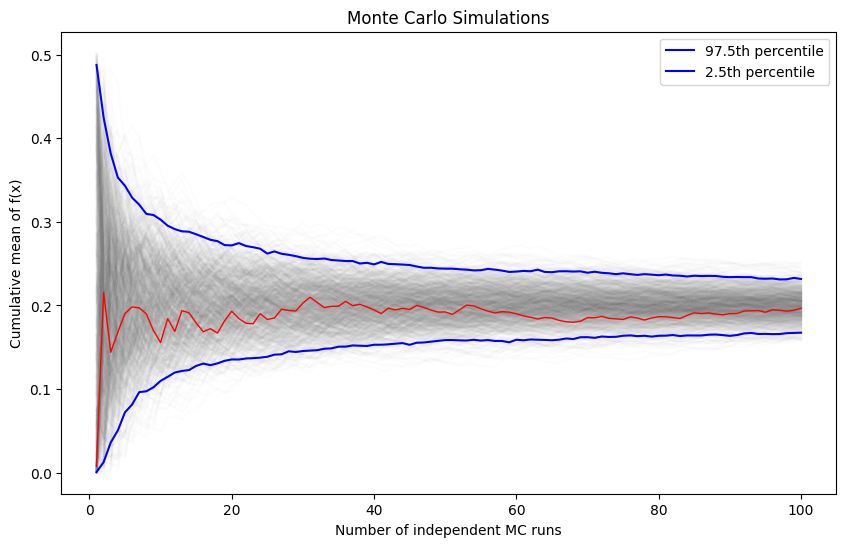
Importance Sampling¶
Suppose we want to evaluate the expectation of a function under a probability distribution :
If sampling directly from is difficult, we can instead introduce an alternative distribution — one that is easier to sample from — and rewrite the integral as:
This reformulation allows us to draw samples , and weight them by the importance ratio . The expectation can then be estimated using:
This is the essence of importance sampling: reweighting samples from an easier distribution to approximate expectations under a more complex one.
import matplotlib as mpl
# Define the unnormalized target distribution: p(y) ∝ exp(-y^4 + y^2)
def p(y):
return np.exp(-y**4 + y**2)
# Uniform proposal over [a, b]
a, b = -5, 5
def q_uniform(y):
return np.ones_like(y) / (b - a)
# Gaussian proposal centered at mode of p(y)
mu_q, sigma_q = 0.0, 1.0
def q_better(y):
return (1 / (np.sqrt(2 * np.pi) * sigma_q)) * np.exp(-(y - mu_q)**2 / (2 * sigma_q**2))
# Estimate normalizing constant Z = ∫ p(y) dy at various sample sizes
sample_sizes = np.logspace(2, 4.5, num=20, dtype=int)
uniform_estimates, better_estimates = [], []
for N in sample_sizes:
y_u = np.random.uniform(low=a, high=b, size=N)
uniform_estimates.append(np.mean(p(y_u) / q_uniform(y_u)))
y_g = np.random.normal(loc=mu_q, scale=sigma_q, size=N)
better_estimates.append(np.mean(p(y_g) / q_better(y_g)))
# Convergence plot
plt.figure(figsize=(10, 5))
plt.plot(sample_sizes, uniform_estimates, 'o-', color='mediumslateblue', label='Uniform proposal')
plt.plot(sample_sizes, better_estimates, 's-', color='seagreen', label='Gaussian proposal')
plt.axhline(y=better_estimates[-1], ls='--', color='gray', label='Reference')
plt.xscale('log')
plt.xlabel('Sample size (log scale)')
plt.ylabel('Importance sampling estimate of Z')
plt.title('Convergence: Gaussian proposal wins because it matches the target shape')
plt.legend()
plt.grid(True, which='both', ls=':', alpha=0.7)
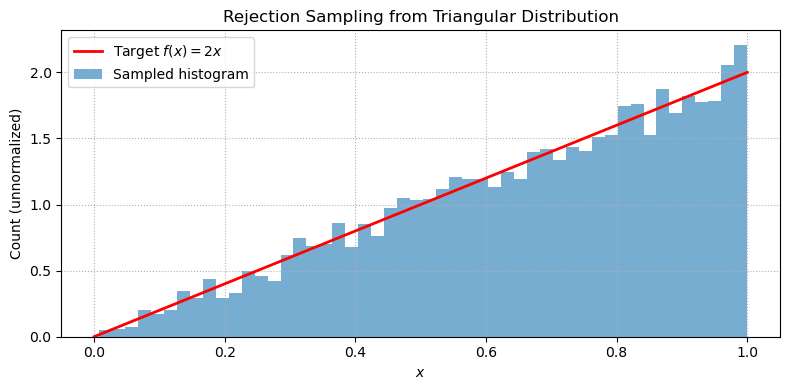
plt.tight_layout();Rejection Sampling¶
Rejection sampling turns uniformly distributed random numbers into samples from a target distribution.
The key idea: draw a candidate uniformly and a height uniformly up to a bounding envelope. If , accept ; otherwise reject. The accepted samples follow the distribution .
Source
import numpy as np
import matplotlib.pyplot as plt
# Target unnormalized PDF: f(x) = 2x on [0, 1]
f = lambda x: 2 * x
Lx, Ly, N = 1, 2, 10000 # Domain, max height, number of samples
# Rejection sampling
x = Lx * np.random.rand(N)
y = Ly * np.random.rand(N)
accepted = x[y <= f(x)]
# Plot result
x_plot = np.linspace(0, 1, 500)
plt.figure(figsize=(8, 4))
plt.plot(x_plot, f(x_plot), 'r-', lw=2, label='Target $f(x) = 2x$')
plt.hist(accepted, bins=50, alpha=0.6, density=True, label='Sampled histogram')
plt.title('Rejection Sampling from Triangular Distribution')
plt.xlabel('$x$')
plt.ylabel('Count (unnormalized)')
plt.legend()
plt.grid(True, linestyle=':')
plt.tight_layout()
plt.show()
Problems¶
Problem 1: MC with importance sampling¶
Evaluate the following integral using Monte Carlo methods:
Start by doing a direct Monte Carlo estimate with uniform sampling on a finite interval .
Try an importance sampling approach using an exponential distribution .
Find the optimal value of that gives the most rapid reduction of variance. [Hint: experiment with different values of and plot the variance vs .]
Problem 2: MC integral of 3D and 6D spheres¶
Generalize the MC code above for computing the volume of 3D and 6D spheres.
The analytical results are known: and . So you can check the statistical error made in the simulations.
Problem 3: Convergence rate of MC¶
Consider the integral .
Estimate using uniform MC with sample sizes .
For each , repeat the estimate 500 times and compute the standard deviation of the estimates.
Plot the standard deviation vs on a log-log scale. Verify that the error decreases as .
How does the prefactor (the variance of on ) compare to your observed intercept?
Problem 4: Rejection sampling from the Boltzmann distribution¶
Use rejection sampling to draw samples from the Boltzmann distribution of a double-well potential:
Use a uniform proposal distribution over with an appropriate bounding constant .
Sample at two temperatures: (low) and (high).
Plot histograms of accepted samples and overlay the exact Boltzmann distribution .
Report the acceptance rate for each temperature. Why does it change?