Quantization of Phase Space in Units of ¶
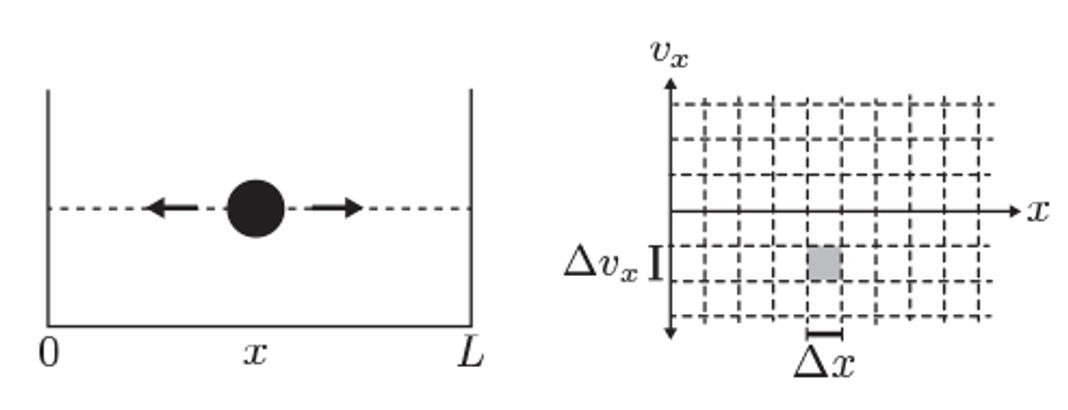
Illustration of phase space for a 1D particle in a box.
In classical mechanics, a system’s state is represented as a point in phase space, with coordinates for each degree of freedom. The number of microstates is, in principle, infinite because both position and momentum can vary continuously.
However, in quantum mechanics, the Heisenberg uncertainty principle imposes a fundamental limit on how precisely position and momentum can be specified:
This implies that we cannot resolve phase space with arbitrary precision. Instead, each distinguishable quantum state occupies a minimum phase space volume on the order of:
For a system with degrees of freedom (e.g for particles in three dimensions), the smallest resolvable cell in phase space has volume:
To compute the number of accessible microstates in a given region of phase space, we divide the classical phase space volume by the smallest quantum volume
Since N particles are indistingusihable and classical mechanics has no such concept we have to manually divide numer of microstates by
An explicit expression for number of microstates for a given energy is written by using delta function which counts all microstates in phase-space where
Phase-space of a harmonic oscillator
= position, = momentum, = mass, = angular frequency
This Hamiltonian gives the total energy of a particle undergoing simple harmonic motion.
Setting , we get the constant-energy curve:
This is the equation of an ellipse in the phase space, with:
Semi-major axis: (in )
Semi-minor axis: (in )
Compute Area of the Ellipse (Classical Action)
Plugging in quantum mechanical expression of quantized energies of harmonic oscillator
This area corresponds to the classical action integral , which in semiclassical quantization becomes:
Source
import numpy as np
import matplotlib.pyplot as plt
hbar = 1.0
m = 1.0
omega = 1.0
n_values = [0, 1, 2, 5, 10]
colors = plt.cm.viridis(np.linspace(0, 1, len(n_values)))
fig, ax = plt.subplots(figsize=(5, 5))
theta = np.linspace(0, 2*np.pi, 500)
for n, c in zip(n_values, colors):
E_n = hbar * omega * (n + 0.5)
a = np.sqrt(2 * E_n / (m * omega**2)) # x semi-axis
b = np.sqrt(2 * m * E_n) # p semi-axis
ax.plot(a * np.cos(theta), b * np.sin(theta), color=c, label=f"$n={n}$")
ax.set_xlabel("Position $x$")
ax.set_ylabel("Momentum $p$")
ax.set_title("Harmonic oscillator: constant-energy ellipses\n"
r"Area $= \pi ab = 2\pi E/\omega = (n+\frac{1}{2})\,h$")
ax.set_aspect('equal')
ax.legend(loc='upper right', fontsize=8)
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()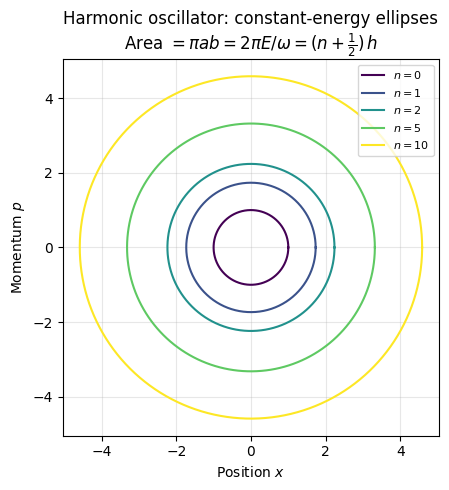
Computing Z via classical mechanics¶
In quantum mechanics counting microstates is easy, we have discrete energies and could simply sum over microstates to compute partition functions
Counting microstates in classical mechanics is therefore done by discretizing phase space for N particle system into smallest unit boxes of size .
Once agan to correct classical mechanics for “double counting” of microstates if we have N indistinguishable “particles” with a factor of
The power and utility of NVT: The non-interacting system¶
For the independent particle system, the energy of each particle enter the total energy of the system additively.
The exponential factor in the partition function allows decoupling particle contributions . This means that the partition function can be written as the product of the partition functions of individual particles!
Distinguishable states:
Indistinguishable states:
Maxwell-Boltzmann distribution¶
The momentum distribution factorizes from the position distribution and is always Gaussian — independent of the interaction potential. This leads to the universal Maxwell-Boltzmann speed distribution for any equilibrium system.
Since the kinetic and potential energy functions depend separately on momenta and positions, the total probability distribution factorizes into a product of position and momentum distributions:
This separation leads to:
The integral with the position-dependent potential for interacting particle system is impossible to evaluate except by simulations or numerical techniques
The momentum-dependent part splits into individual integrals for every particle and is always a simple quadratic function, making it analytically tractable:
The momentum (or velocity) distribution follows the Maxwell-Boltzmann distribution in all equilibrium systems, regardless of the complexity of the molecular interactions.
Maxwell-Boltzmann Velocity Distribution: Full Derivation
Separation of variables
In equilibrium, the distribution of velocities in the , , and directions are independent and identical, because there’s no preferred direction in an ideal gas. Therefore, we can write the joint probability distribution as:
Furthermore, the probability of finding a particle with velocity components between , , and is:
Form of the velocity component distribution
The system is in thermal equilibrium, so the probability of a particle having a certain velocity corresponds to the Boltzmann factor for its kinetic energy:
For a single particle of mass , the kinetic energy is:
So we postulate:
Using the independence of the components:
We require that:
This is a standard Gaussian integral of form where and therefore we write down:
Distribution of speeds
Now, let’s consider the speed .
The probability density for the speed (regardless of direction) is obtained by converting to spherical coordinates in velocity space. The volume element in velocity space becomes:
Integrating over angles gives a factor of :
Substitute in the expressions for , , and , and recall that the total exponent is:
Therefore, the speed distribution is:
This is the Maxwell-Boltzmann speed distribution. It gives the probability density that a particle in a gas has speed .
Source
import numpy as np
import matplotlib.pyplot as plt
k_B = 1.38e-23 # J/K
m = 4.65e-26 # kg (nitrogen molecule, approximate)
T = 300 # K
v = np.linspace(0, 2000, 1000)
def f_MB(v, m, T):
return 4*np.pi * v**2 * (m / (2*np.pi*k_B*T))**(3/2) * np.exp(-m*v**2 / (2*k_B*T))
v_mp = np.sqrt(2 * k_B*T / m)
v_avg = np.sqrt(8 * k_B*T / (np.pi*m))
v_rms = np.sqrt(3 * k_B*T / m)
fig, ax = plt.subplots(figsize=(7, 4))
ax.plot(v, f_MB(v, m, T), color='black', label="Maxwell-Boltzmann")
ax.axvline(v_mp, color='C0', ls='--', label=f"$v_{{mp}}$ = {v_mp:.0f} m/s")
ax.axvline(v_avg, color='C2', ls='--', label=f"$\\langle v \\rangle$ = {v_avg:.0f} m/s")
ax.axvline(v_rms, color='C3', ls='--', label=f"$v_{{rms}}$ = {v_rms:.0f} m/s")
ax.set_xlabel("Speed (m/s)")
ax.set_ylabel("Probability density")
ax.set_title("Maxwell-Boltzmann speed distribution — $N_2$ at 300 K")
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()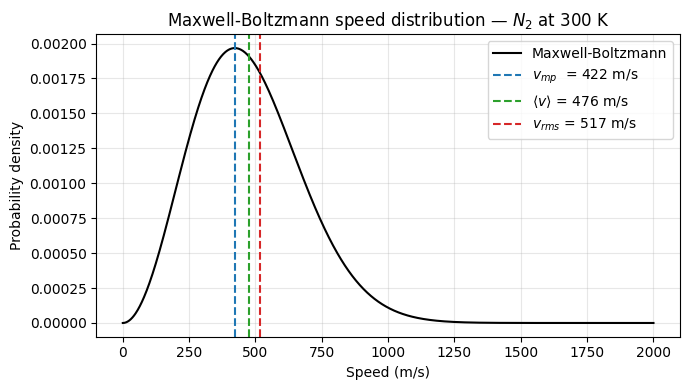
Ideal Gas¶
A useful approximation is to consider the limit of a very dilute gas, known as an ideal gas, where interactions between particles are neglected ().
Evaluating the partition function for a single atom of an ideal gas:
On the validity of the classical approximation
The classical approximation is valid when the average interatomic spacing is much larger than the thermal de Broglie wavelength:
This ensures that wavefunction overlap between particles is negligible, so quantum exchange statistics can be ignored and particles obey Maxwell-Boltzmann statistics for their translational motion.
We can also calculate entropy and other thermodynamic quantities:
Equipartition Theorem¶
Consider a classical system with a Hamiltonian containing a quadratic degree of freedom , such that:
The canonical partition function for this degree of freedom is:
The average energy is:
Examples¶
Monatomic ideal gas: 3 translational degrees →
Diatomic molecule at high : 3 translational + 2 rotational + active vibrational modes
→
Partition functions for non-interacting molecules¶
Total energy and partition function for a molecule with separable degrees of freedom:
Translational degrees of freedom: particle in a box¶
For a particle in a 3D box of volume , the energy levels are:
At high temperatures (many levels populated), we replace the discrete sum with an integral:
For indistinguishable molecules:
Vibrational degrees of freedom: harmonic oscillator¶
Energy levels: . Partition function for one mode:
Average energy for independent oscillators:
Low T (): — zero-point energy only.
High T (): — classical equipartition recovered.
Rotational degrees of freedom: rigid rotor¶
For a linear molecule (diatomic rotor), the rotational energy levels are:
At high temperatures (), the sum becomes an integral:
where is the rotational temperature.
Source
import numpy as np
import matplotlib.pyplot as plt
k_B = 1.0 # work in units of k_B
hbar_omega = 1.0 # energy quantum (sets the scale)
T_vals = np.linspace(0.05, 5.0, 500) # in units of ℏω/k_B
x = hbar_omega / T_vals # dimensionless ℏω/k_BT
C_v = k_B * x**2 * np.exp(x) / (np.exp(x) - 1)**2
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(T_vals, C_v, color='darkorange', lw=2)
ax.axhline(1, color='gray', ls='--', lw=1, label=r"Classical limit $k_B$")
ax.set_xlabel(r"$k_B T \,/\, \hbar\omega$")
ax.set_ylabel(r"$C_V \,/\, k_B$")
ax.set_title("Heat capacity of a quantum harmonic oscillator")
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()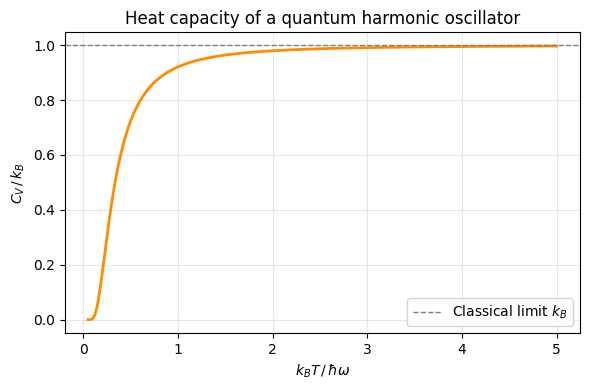
On heat capacity of harmonic-oscillators¶
Einstein Model of solids¶
A simple model for solids due to Einstein describes it as independent harmonic oscillators representing localized atomic vibrations.
The average energy of a quantum harmonic oscillator is:
The heat capacity is the derivative:
Key predictions of Einstein model of solids¶
At low T: — the oscillator is frozen in its ground state.
At high T: — classical equipartition is recovered.
Debye Model of Solids¶
The Debye model improves upon Einstein’s and matches experimental data much better than the Einstein model, especially at low temperatures.
Main distinguishing features of Debye model are:
Treating atomic vibrations as a continuum of phonon modes.
Including a full spectrum of frequencies up to a Debye cutoff .
Predicting the correct low-temperature behavior of solids.
Behavior of Debye Model¶
Low T: — from long-wavelength (acoustic) phonons.
High T: — classical limit.
Source
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import quad
k_B = 1.0 # work in units of k_B
# --- Einstein model: 3 modes per atom ---
def einstein_cv(T, T_E):
x = T_E / T
return 3 * k_B * x**2 * np.exp(x) / (np.exp(x) - 1)**2
# --- Debye model: integrates over full phonon spectrum ---
def debye_cv(T, T_D):
x_D = T_D / T
integrand = lambda x: x**4 * np.exp(x) / (np.exp(x) - 1)**2
val, _ = quad(integrand, 1e-10, x_D)
return 9 * k_B * (T / T_D)**3 * val
T_D = 1.0
T_E = 0.75 * T_D # empirical best-fit ratio
T_vals = np.linspace(0.01, 3.0, 300) # in units of T_D
cv_einstein = [einstein_cv(T, T_E) for T in T_vals]
cv_debye = [debye_cv(T, T_D) for T in T_vals]
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
for ax, xscale, title in zip(axes, ['linear', 'log'], ['Linear scale', 'Log scale (low-T behavior)']):
ax.plot(T_vals, cv_einstein, color='darkorange', lw=2, label=r'Einstein ($\theta_E = 0.75\, T_D$)')
ax.plot(T_vals, cv_debye, color='steelblue', lw=2, label=r'Debye ($\theta_D = T_D$)')
ax.axhline(3, color='gray', ls='--', lw=1.2, label=r'Dulong–Petit limit ($3k_B$)')
ax.set_xscale(xscale)
ax.set_xlabel('$T / T_D$')
ax.set_ylabel('$C_V / k_B$')
ax.set_title(title)
ax.set_ylim(0, 3.5)
ax.legend()
ax.grid(True, alpha=0.4)
fig.suptitle('Einstein vs Debye Model: Heat Capacity of Solids', fontsize=13, fontweight='bold')
plt.tight_layout()
plt.show()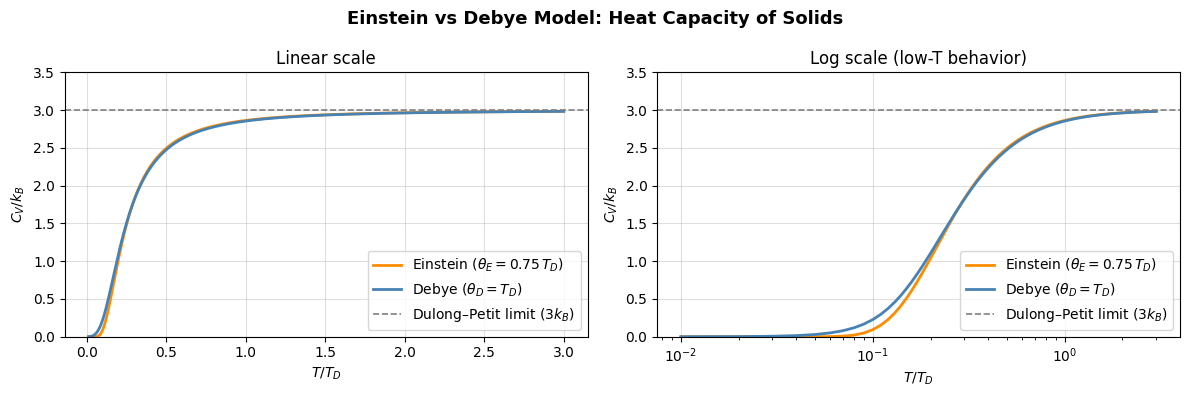
Problems¶
Problem-1 1D energy profiles¶
Given an energy function with constants
Compute free energy difference between minima at temperature
Compute free energy around minima of size
Problem-2 Maxwell-Boltzmann¶
You are tasked with analyzing the behavior of non-interacting nitrogen gas molecules described by the Maxwell-Boltzmann distribution:
Where:
is the probability density for molecular speed
is the mass of a nitrogen molecule
(Boltzmann constant)
is the temperature in Kelvin
Step 1: Plot the Maxwell-Boltzmann Speed Distribution
At room temperature (T = 300, 400, 500, 600 K), plot the distribution for speeds ranging from 0 to 2000 m/s.
Step 2: Identify Most Probable, Average, and RMS Speeds
Compute and print:
The most probable speed
The mean speed
The root mean square (RMS) speed
Plot vertical lines on your plot from Step 1 to show these three characteristic speeds.
Step 3: Speed Threshold Analysis
Use numerical integration to calculate the fraction of molecules that have speeds:
(a) Greater than 1000 m/s
(b) Between 500 and 1000 m/s
Hint: Use
scipy.integrate.quadto numerically integrate the Maxwell-Boltzmann distribution.
Problem-3 Elastic Collisions¶
In this problem need to modify the code to add elastic collisions to the ideal gas simulations
Problem-4 Elementary derivation of Boltzmann’s distribution¶
Let us do an elementary derivation of the Boltzmann distribution showing that when a macroscopic system is in equilibrium and coupled to a heat bath at temperature we have a universal dependence of probability for finding the system at different energies:
The essence of the derivation is this. Consider a vertical column of gas somewhere in the mountains.
On one hand we have gravitational force which acts on a column between with cross section .
On the other hand we have pressure balance which keeps the molecules from dropping to the ground.
This means that we have a steady density of molecules at each height for a fixed . Write down this balance of forces (gravitational vs pressure) and show how density at , is related to density at , .
Tip: you may use for pressure and for the gravitational potential energy.
Problem-5 2D dipoles on a lattice¶
Consider a 2D square lattice with lattice points. On each point we have a magnetic moment that can point in four possible directions: . Along the axis, the dipole has energy and along the axis .
Dipoles are not interacting with each other and we also ignore kinetic energy of dipoles since they are fixed at lattice positions.
Write down the partition function for this system
Compute the average energy.
Compute entropy per dipole . Evaluate the difference . Can you see a link with the number of arrangements of dipoles?
Compute the microcanonical partition function
Show that we get the same entropy expression from both the and ensembles.